FLASH: Fast Landmark Aligned Spherical Harmonic Parameterization for Genus-0 Closed Brain Surfaces
Surface registration between cortical surfaces is crucial in medical imaging for performing systematic comparisons between brains. Landmark-matching registration that matches anatomical features, called the sulcal landmarks, is often required to obtain a meaningful 1-1 correspondence between brain surfaces. This is commonly done by parameterizing the surface onto a simple parameter domain, such as the unit sphere, in which the sulcal landmarks are consistently aligned. Landmark-matching surface registration can then be obtained from the landmark aligned parameterizations. For genus-0 closed brain surfaces, the optimized spherical harmonic parameterization, which aligns landmarks to consistent locations on the sphere, has been widely used. This approach is limited by the loss of bijectivity under large deformations and the slow computation. In this paper, we propose FLASH, a fast algorithm to compute the optimized spherical harmonic parameterization with consistent landmark alignment. This is achieved by formulating the optimization problem to the extended complex plane and thereby linearizing the problem. Errors introduced near the pole are corrected using quasi-conformal theories. Also, by adjusting the Beltrami differential of the mapping, a diffeomorphic (1-1, onto) spherical parameterization can be effectively obtained. The proposed algorithm has been tested on 38 human brain surfaces. Experimental results demonstrate that the computation of the landmark aligned spherical harmonic parameterization is significantly accelerated using the proposed algorithm.
Example 1:
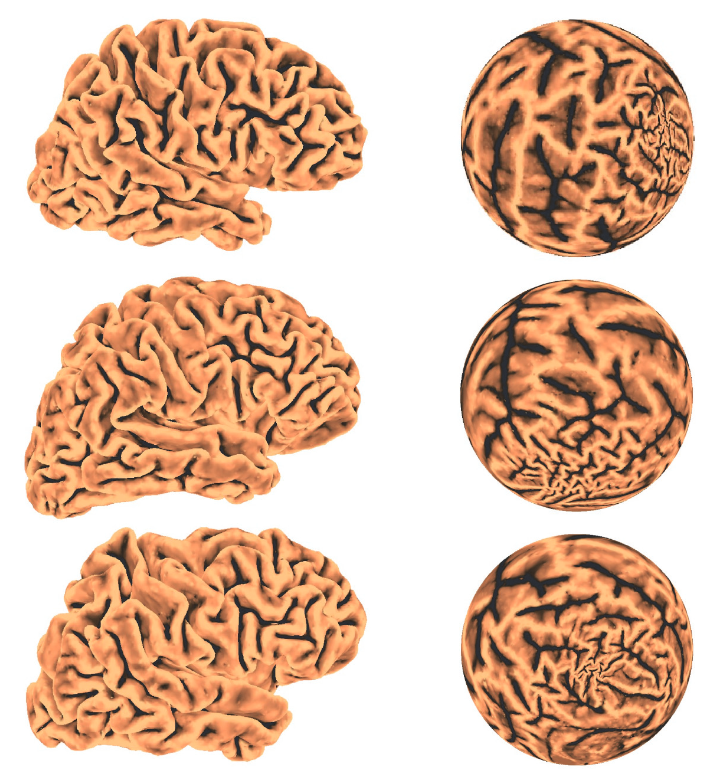
Spherical conformal parameterizations using our proposed algorithm. Left: Three input braincortical surfaces. Right: The spherical conformal parameterizations obtained by our proposed algorithm.
Example 2:
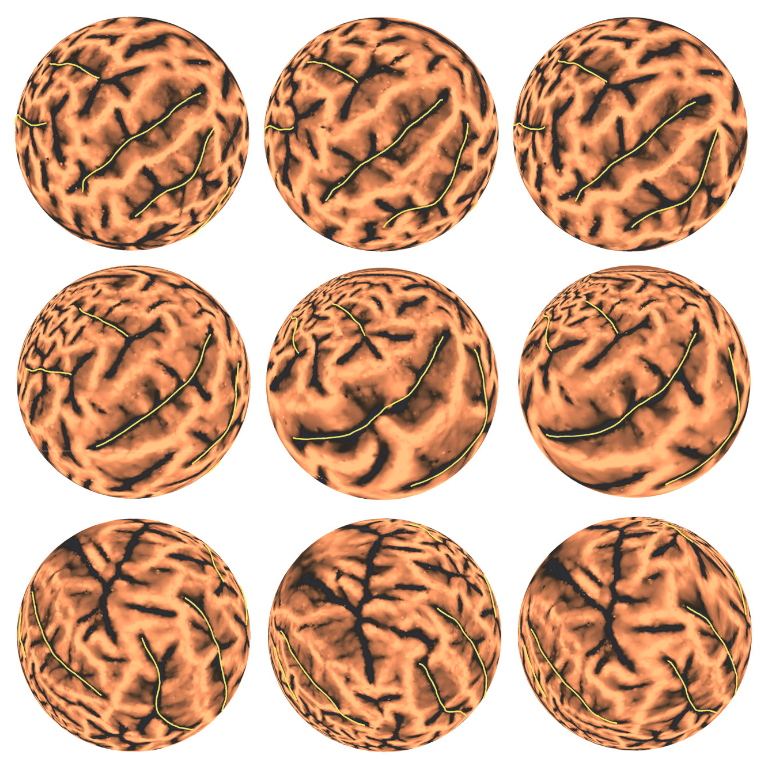

Landmark aligned spherical harmonic parameterizations using FLASH. One set of experimental results is shown in each row. The spherical conformal parameterizations of the source brains and the target brains are shown, respectively, in the first and the second column. The corresponding landmark aligned spherical harmonic parameterizations are given in the third column.
Example 3:
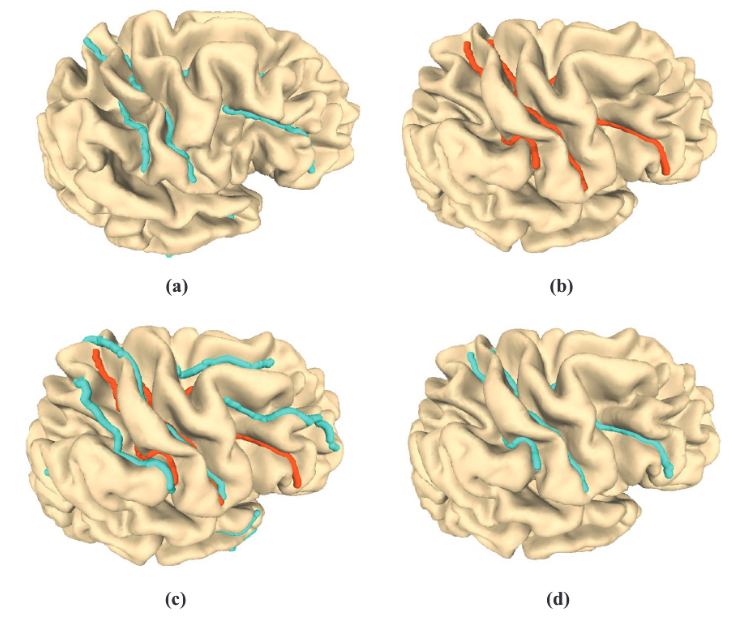
Two cortical surfaces and the registration results. The sulcal landmarks are highlighted. (a) and (b) show the source cortical surface and the target cortical surface, respectively. (c) shows the conformal registration without any landmark constraints. One can observe that the landmark curves are not matched. (d) shows the registration obtained using FLASH.