Shape Analysis of Partial Surfaces via Inconsistent Surface Mapping technique
Project Description:
We
develop a framework for shape analysis of partial surfaces using
inconsistent surface mapping technqiue. Traditional landmark-based
geometric morphometrics methods suffer from the limited degrees of
freedom, while most of the more advanced non-rigid surface mapping
methods rely on a strong assumption of the global consistency of two
surfaces. From a practical point of view, given two anatomical surfaces
with prominent feature landmarks, it is more desirable to have a method
that automatically detects the most relevant parts of the two surfaces
and finds the optimal landmark-matching alignment between those parts,
without assuming any global 1-1 correspondence between the two partial
surfaces. Our method is capable of solving this problem using an
inconsistent surface mapping technique based on quasi-conformal theory.
It further enables us to quantify the dissimilarity of two shapes using
quasi-conformal distortion and differences in mean and Gaussian
curvatures, thereby providing a natural way for shape classification.
Experiments on Platyrrhine molars demonstrate the effectiveness of our
method and shed light on the interplay between function and shape in
nature.
Publication:
- P.T. Choi, S. Di, L.M. Lui, Shape Analysis of Partial Surfaces via Inconsistent Surface Mapping technique (2020)
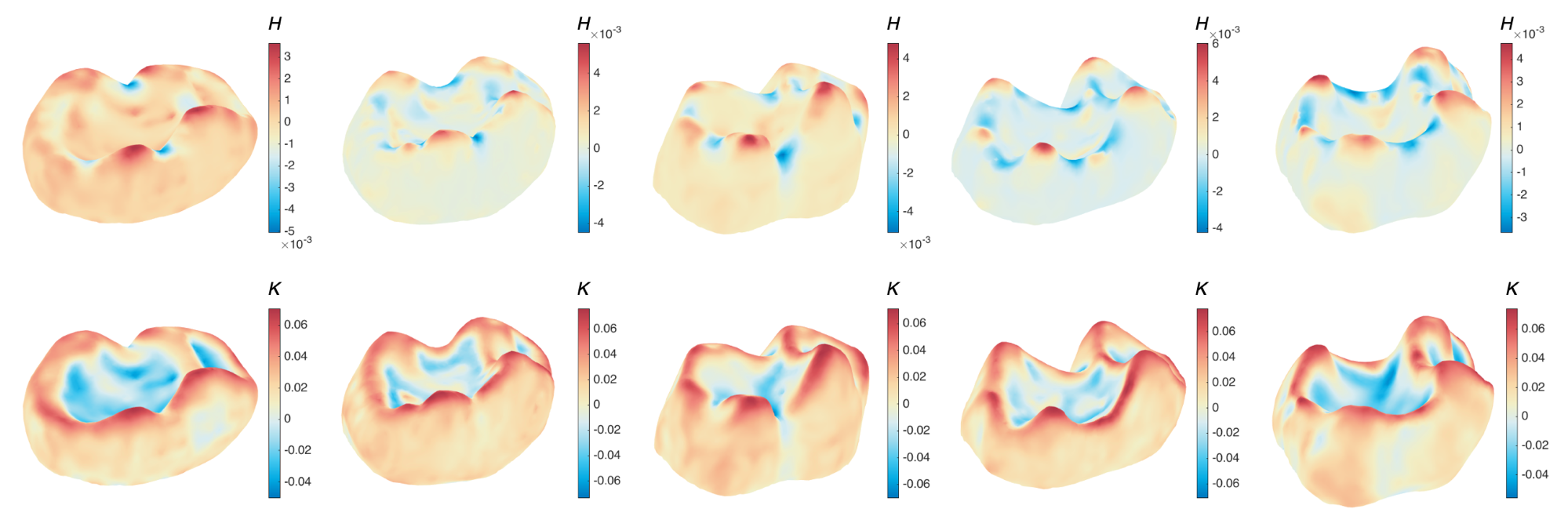
Five example tooth surfaces in the mammalian molar dataset, color-coded by the mean curvature H and the Gaussian curvature K. Each column corresponds to one tooth surface..
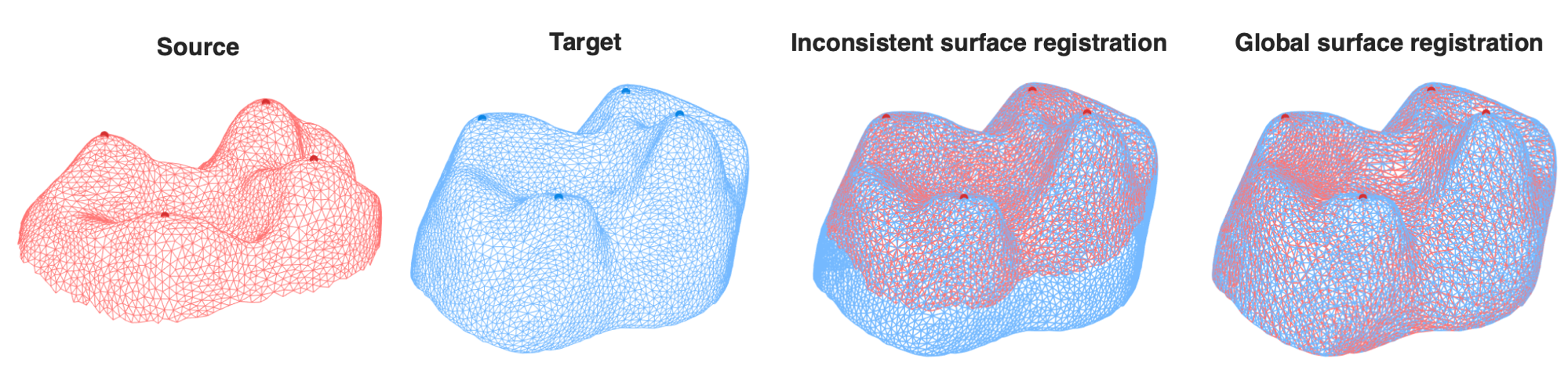

A comparison between our inconsistent surface registration approach and the globallandmark-matching quasi-conformal Teichmuller mapping method. Here, the source mesh(leftmost) and the target mesh (second left) are inconsistently segmented. Our inconsistent surfaceregistration approach successfully identifies and registers the common regions of the two meshes(second right). On the contrary, the landmark-matching Teichmuller mapping method constructsa global 1-1 correspondence between the two meshes including the inconsistent parts, thereby leadingto an overall misalignment with a large distortion (rightmost).
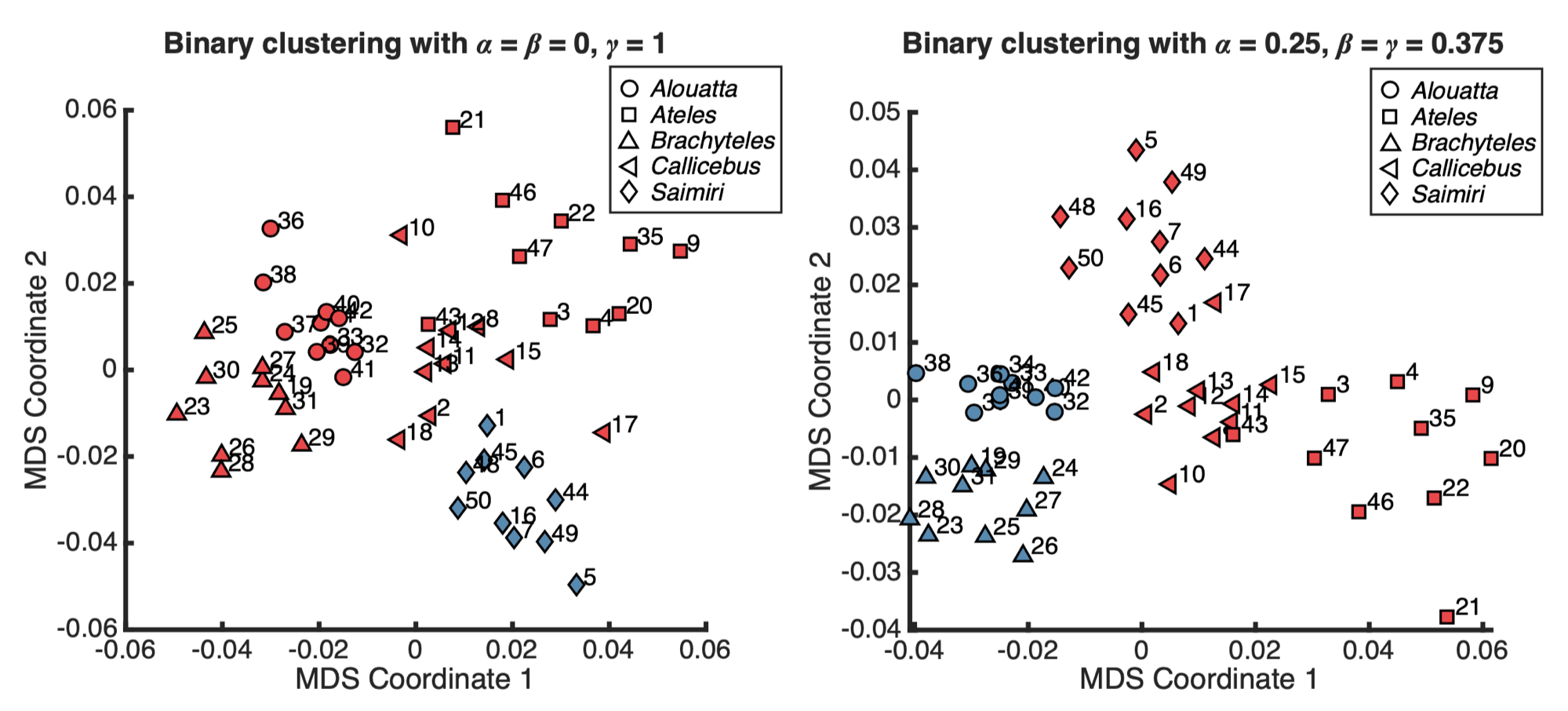
The binary clustering results produced by the hierarchical clustering method using our proposed dissimilarity measure, visualized on the multidimensional scaling (MDS) plane. The specimens from different genera are represented using different markers, and the colors of the markers represent the clustering result. The numerical label of each node corresponds to the specimen ID. For the result shown on the left, the parameters (α,β,γ) = (0,0,1) areused for constructing the dissimilarity matrixD. One of the resulting clusters consists of all Saimiri specimens (insectivores), while the other one consists of all non-insectivores. For the result shown on the right, the parameters (α,β,γ) = (0.25,0.375,0.375) are used for constructing D. One of the resulting clusters consists of all Alouatta and Brachyteles specimens (folivores), while the other one consists of all non-folivores.
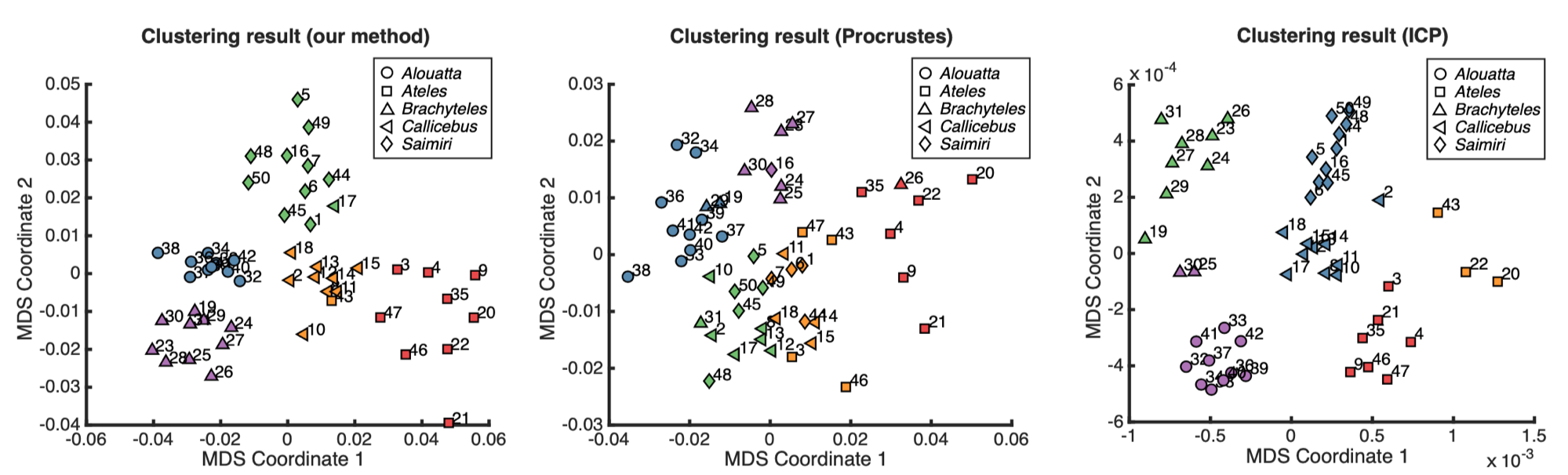
The clustering results produced by the k-means clustering method with $k = 5$ using different dissimilarity measures, visualized on the multidimensional scaling (MDS) plane. The specimens from different genera are represented using different markers, and the colors of the markers represent the clustering result. The numerical label of each node corresponds to the specimen ID. Left: the clustering result obtained from our method. Middle: the clustering result obtained from the Procrustes superimposition method. Right: the clustering result obtained from the Iterative Closest Point (ICP) method.