Automatics sulcal landmark tracking on the brain cortical surface
Project Description:
One important problem in the human brain mapping research
is to locate the important anatomical features. Anatomical features on thecortical
surface are usually represented by landmark curves, called
sulci/gyri curves. These landmark curves are important information for neuroscientists
to study brain disease and to match different cortical surfaces. Manual labelling
of these landmark curves is
time-consuming, especially when large set of data has to be analyzed. In
this paper, we present algorithms to automatically detect and match landmark
curves on cortical surfaces to get an optimized brain conformal parametrization.
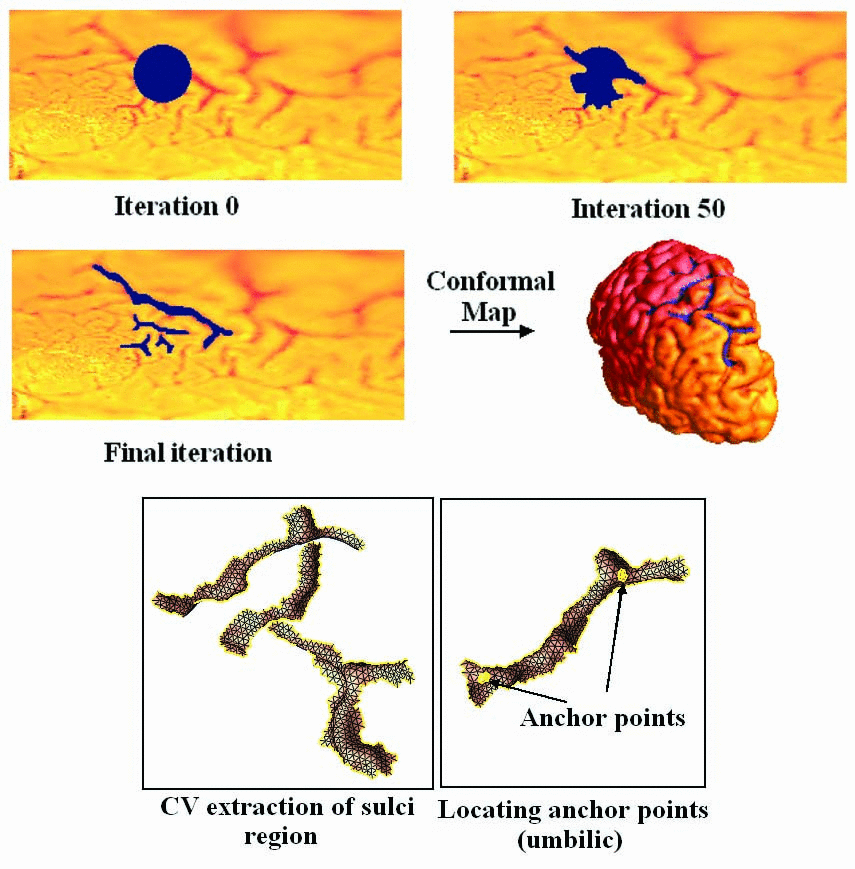
First, we propose an algorithm to obtain a hypothesized landmark region/curves
using the
Chan-Vese segmentation method, which solves a Partial Differential Equation
(PDE) on a manifold with global conformal parameterization. This is done by
segmentating the high mean curvature region. Second,
we propose an automatic landmark curve tracing method based on the principal
directions of the local Weingarten matrix. Based on the global conformal parametrization
of a cortical surface, our method
adjusts the landmark curves iteratively on the spherical or rectangular
parameter domain of the cortical surface along its principal direction field,
using umbilic points of the surface as anchors. The landmark curves can then
be mapped back onto the cortical surface. Experimental results show that
the landmark curves detected by our algorithm closely resemble these manually
labeled curves. Next, we applied these automatically labeled landmark curves
to generate an optimized conformal parametrization of the cortical
surface, in the sense that homologous features across subjects are caused
to lie at the same parameter locations in a conformal grid. Experimental results
show that our method can effectively help in
automatically matching cortical surfaces across subjects.
Publication:
- L.M. Lui, Y. Wang, T.F. Chan, and P.M. Thompson, "Brain Anatomical Feature Detection by Solving Partial Differential Equations on General Manifolds", Discrete and Continuous Dynamical Systems B, 7(3), May 2007, pp. 605-618
- L. M. Lui, Y. Wang, T.F. Chan and P.M. Thompson, "Automatic Landmark Tracking and Its Application to the Optimization of Brain Conformal Mapping", accepted by IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR), New York, NY, Jun. 2006
- L. M. Lui, Y. Wang, T.F. Chan and P.M. Thompson, "Automatic Landmark Tracking Applied to Optimize Brain Conformal Mapping", IEEE International Symposium on Biomedical Imaging: From Nano to Macro (ISBI), Arlington, VA, Apr. 2006, pp. 205-208
Result: